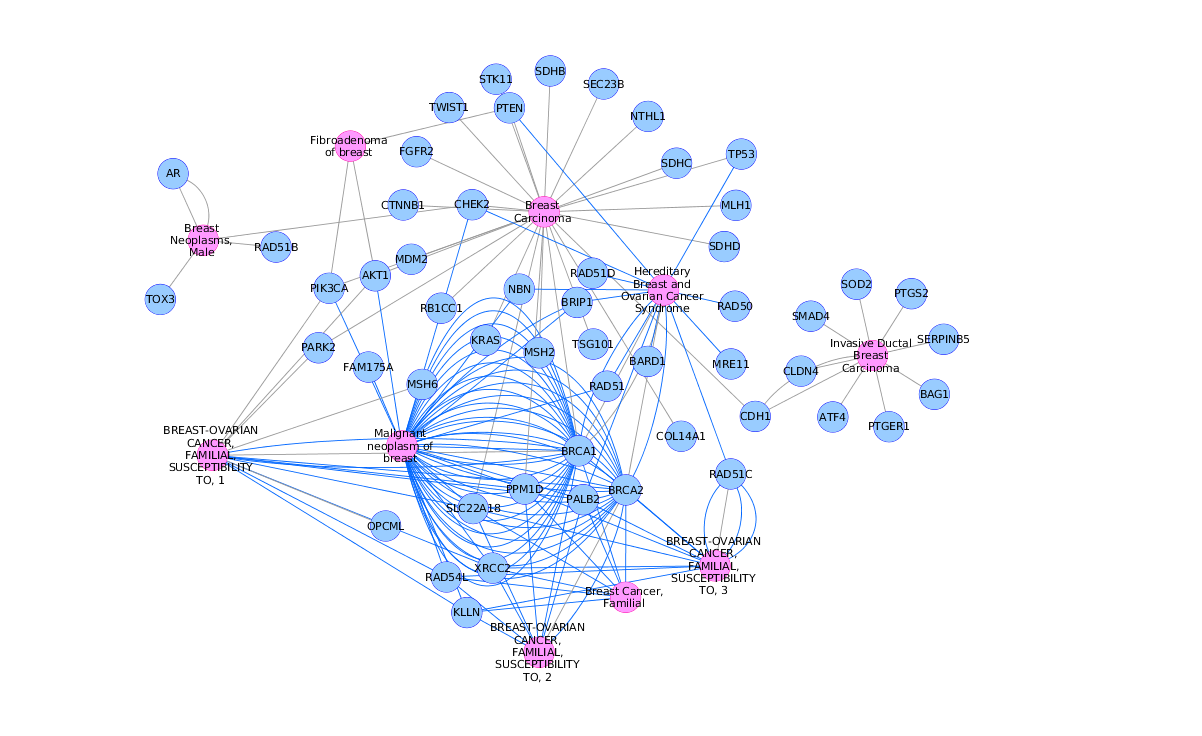
Use the network manager to easily organize multiple networks.Zoom in/out and pan for browsing the network.A variety of layout algorithms are available, including cyclic and spring-embedded layouts. according to user-configurable colors and visualization schemes. 
Expression data can be mapped to node color, label, border thickness, or border color, etc. View a superposition of gene expression ratios and p-values on the network.Customize network data display using powerful visual styles.Cytoscape session file includes networks, attributes (for node/edge/network), desktop states (selected/hidden nodes and edges, window sizes), properties, and visual styles. Load and save state of the cytoscape session in a cytoscape session (.cys) file.Directly import GO terms and annotations from OBO and Gene Association files.Import gene functional annotations from the Gene Ontology (GO) and KEGG databases.For example, input a set of custom annotation terms for your proteins, create a set of confidence values for your protein–protein interactions. Load and save arbitrary attributes on nodes and edges.Input mRNA expression profiles from tab- or space-delimited text files.Load and save networks and node/edge attributes in an XML document format called XGMML (eXtensible Graph Markup and Modeling Language).Load and save previously-constructed interaction networks in GML format (Graph Modelling Language).User-defined interaction types are also supported. For yeast and other model organisms, large sources of pairwise interactions are available through the BIND and TRANSFAC databases. Input and construct molecular interaction networks from raw interaction files (SIF format) containing lists of protein-protein and/or protein–DNA interaction pairs.Plugins are developed by core developers and the greater user community. A vital aspect of the software architecture of Cytoscape is the use of plugins for specialized features. Cytoscape can visualize and analyze network graphs of any kind involving nodes and edges (e.g., social networks). While Cytoscape is most commonly used for biological research applications, it is agnostic in terms of usage. Thank you in advance for your cooperation.Yeast Protein–protein/Protein–DNA interaction network visualized by Cytoscape. A no-show will also limit your ability to book our events in the future. While we do not charge a fee to attend this event, these programs would not be sustainable and available to all wanting to attend, unless all registrants abide by the 48 hour cancellation notice policy. He has given numerous presentations on using and extending Cytoscape and is a Cytoscape core developer as well as the developer of over a dozen Cytoscape apps, including chemViz, structureViz, clusterMaker, and cddApp. Scooter is the executive director of the Resource for Biocomputing, Visualization, and Informatics at UCSF, the “roving engineer” for Cytoscape, and an adjunct assistant professor of Pharmaceutical Chemistry at UCSF. He has been a contributing member to Cytoscape since 2006 and has led numerous Cytoscape and Network Biology workshops and mentoring programs over the past 10 years. Alex is the executive director of the National Resource for Network Biology, the vice president of the Cytoscape Consortium, and associate director of bioinformatics at Gladstone Institutes. PresentersĪlexander Pico, Gladstone Institutes. Installation instructions will be provided in the weeks preceding the tutorial. Participants are required to bring a laptop with Cytoscape installed. A solid background in biology and active research questions are expected. This tutorial is intended for an audience that has no prior experience with Cytoscape. Know where to find relevant Cytoscape apps and tutorials.Master network layouts and data visualization.Perform basic topological and other network analyses.Understand the major applications of network biology.By the end of tutorial, you should be able to This workshop will provide a general introduction to network biology studies and Cytoscape concepts, including a hands-on session for universal data import and demonstration of a few of the over 300 freely available apps contributed by the Cytoscape developer community. As a free and open source tool, Cytoscape has become the most popular network visualization and analysis tool in the biological sciences. The network perspective on biology brings meaningful context to high-throughput data for exploratory analysis, interpretation, and hypothesis generation. Slides for the workshop are available for download here ! Get introduced to network biology studies and Cytoscape concepts.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
